The Promise and the Gap: Why Chatbot Experimentation Isn't Enough
If you work in healthcare analytics, you've almost certainly experimented with ChatGPT or Claude. Maybe you've asked it to summarise a clinical paper, draft a literature search strategy, or explain a statistical concept. You're not alone: 85% of healthcare organisations have now adopted or explored generative AI, up from 72% just a year ago.
Yet despite this widespread experimentation, most of us remain stuck at the surface — using these powerful models as glorified search engines or writing assistants, asking questions, getting answers, copying and pasting results into our workflows. This is chatbot use, and while it has its place, it barely scratches the surface of what generative AI can do for healthcare analytics.
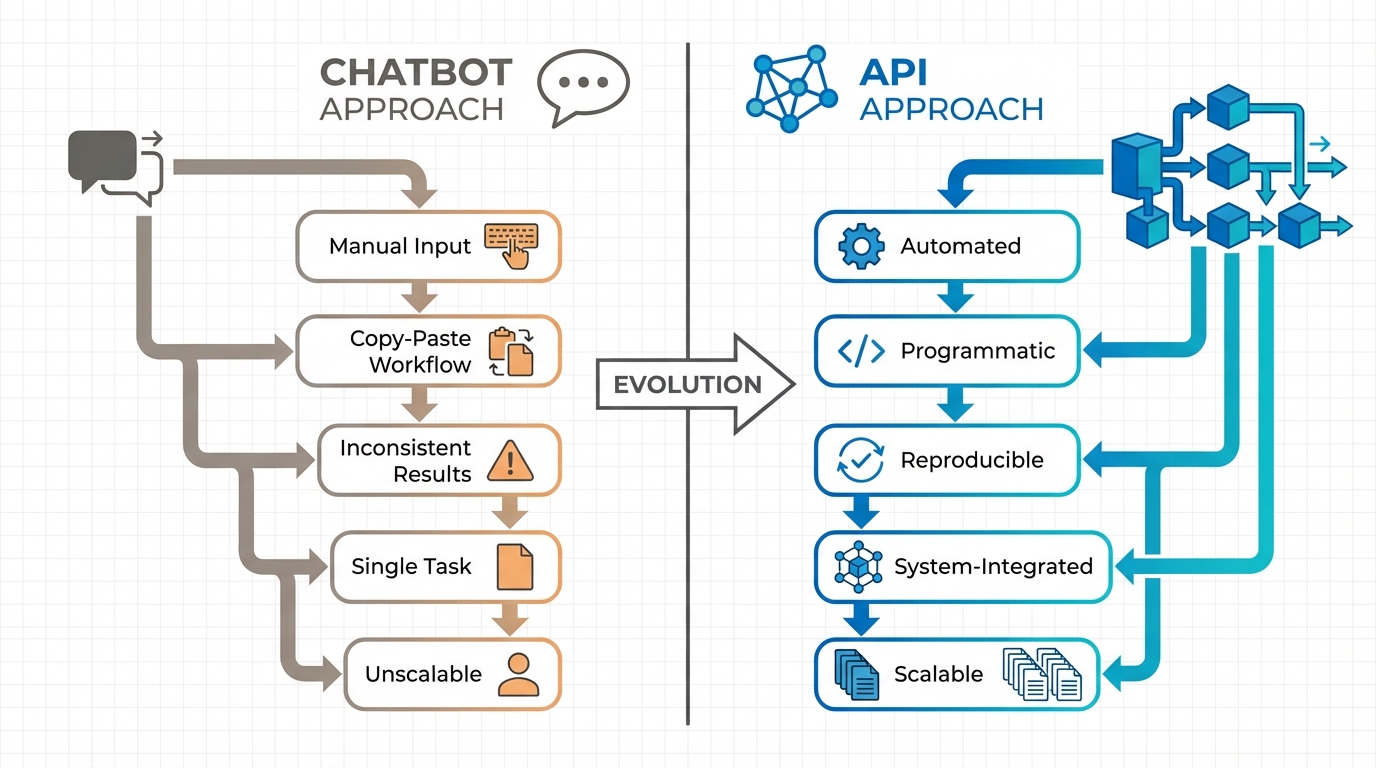
The real transformative potential lies in automating the tedious, time-consuming work that dominates HEOR research. Consider systematic literature reviews: the foundation of evidence synthesis and health technology assessment. Traditional approaches require weeks of manual screening, data extraction, and synthesis. Recent studies demonstrate that properly implemented AI systems can achieve 100% sensitivity in identifying relevant studies while reducing workload by approximately 80%. Collaborative AI approaches using structured extraction workflows now achieve 87–96% accuracy with appropriate quality controls.
These aren't hypothetical promises. Research teams are already using GenAI to reproduce and update entire issues of Cochrane systematic reviews in under two days — work that would traditionally require months. The healthcare AI market reflects this shift: projected growth from $2.17 billion in 2024 to $14.82 billion by 2030 signals that organisations recognise the stakes.
The Gap That Matters
Here's the problem: there's an enormous gap between "playing with ChatGPT" and "building reliable AI-enabled research tools." That gap isn't crossed by crafting better prompts or spending more time in the chat interface. It requires deliberate skill development in five specific, learnable areas — areas that most healthcare analytics professionals have not needed in the past.
The transition doesn't require becoming a software engineer or abandoning domain expertise. It requires adding foundational technical skills that, in the era of AI assistants, are more accessible than ever before. The five skills that changed how I approach every research problem are:
The Five Skills
- Learning Python basics
- Accessing GenAI through APIs rather than chatbots
- Using structured outputs to control AI behaviour
- Thinking in terms of systems rather than single tasks
- Building monitoring into AI workflows
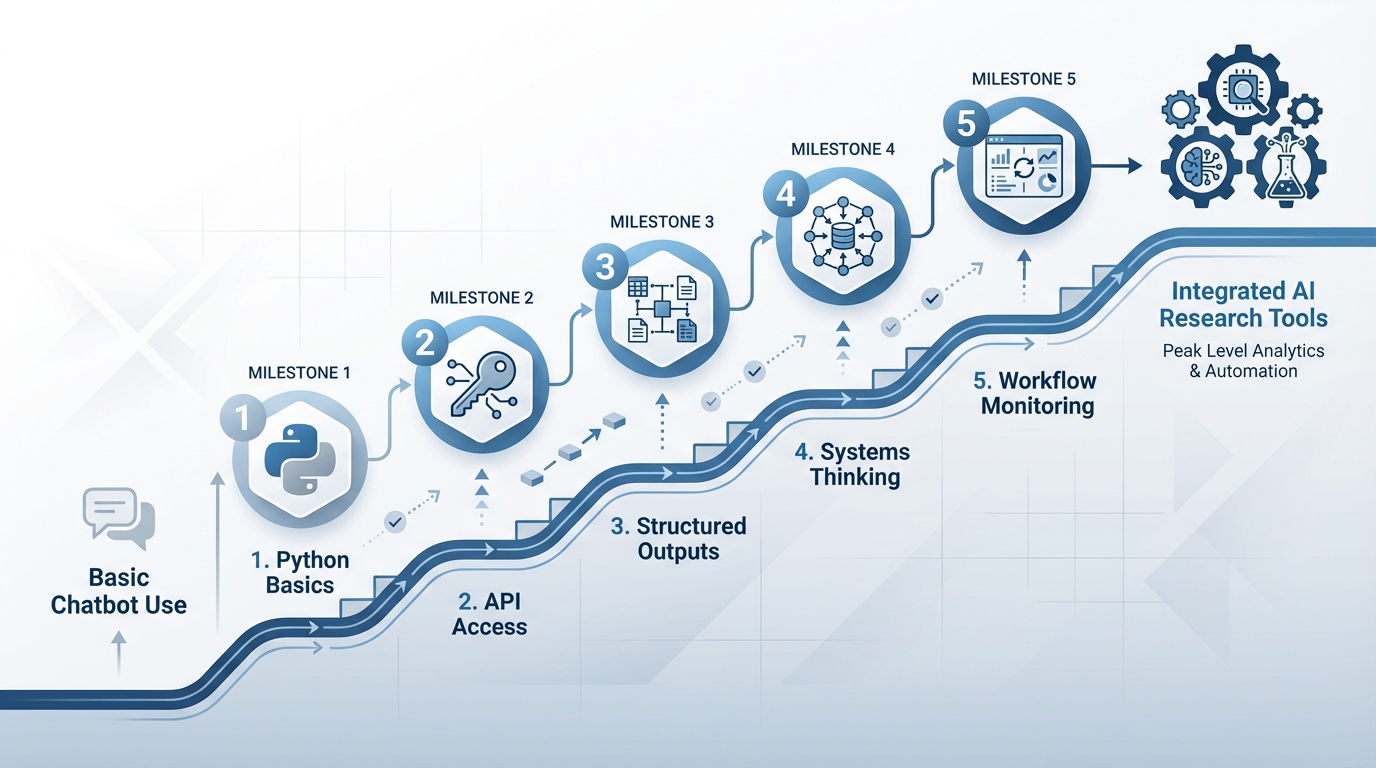
None of these are mysterious. Each is learnable over a few focused weekends. Together, they represent the difference between using AI as an occasional helper and integrating it as a reliable research tool.
Skill 1 & 2: Building the Foundation — Python and API Access
If I told you five years ago that healthcare researchers would need to learn programming to stay competitive, you might have been skeptical. Today, the question isn't whether programming skills matter — it's how much you actually need to know, and whether learning is realistic given everything else on your plate.
Here's what I've learned: you don't need to become a software engineer. You need to understand enough to work intelligently with AI-generated code, to know when something makes sense and when it doesn't, and to connect the pieces that turn isolated AI outputs into functioning research tools.
Why Python, and Why Now
Python has become the de facto language for AI integration because it balances accessibility with professional capability. Unlike statistical software limited to specific workflows, Python lets you combine data processing, API calls to AI models, file handling, and output generation in a single script. More importantly, it's what AI assistants are trained on — when you ask Claude or ChatGPT to write code, they'll almost always produce Python.
The practical implication: effective GenAI applications require combining AI with deterministic code. Consider extracting structured data from 100 clinical trial papers. The AI model handles the intelligent pattern recognition — reading each paper and identifying study design, population size, endpoints, and results. But deterministic code handles everything else: reading files from a folder, sending each to the API with your extraction schema, collecting responses, validating the structure, writing results to a spreadsheet, and logging any errors. You need programming skills to orchestrate this workflow. "Basics, not expertise" is the right target.
From Chatbots to APIs: The Critical Transition
The chatbot web interface is fundamentally limited: paste content into a text box, the model generates a response, and you copy the result somewhere else. This workflow is fine for one-off questions, but it breaks down for systematic research tasks. How do you process 50 documents consistently? How do you ensure the output format matches what your analysis needs? How do you log what happened for quality control?
API access solves this. Instead of conversing with a chatbot through a web interface, you write code that sends information to the model programmatically and receives structured responses. This architectural shift unlocks capabilities impossible in chatbot interfaces: feeding the model proprietary data it has never seen, defining exactly what structure the response should take, building workflows where one AI call's output becomes the next step's input, and logging every input and output for monitoring, debugging, and compliance documentation.
The practical learning curve is gentle. Moving from ChatGPT's web interface to making your first API call takes perhaps an hour. All major models — OpenAI's GPTs, Anthropic's Claude, Google's Gemini — offer API access with comparable capabilities. The choice depends on your use case, but the pattern is consistent: you authenticate with an API key, send a request containing your prompt and any documents or data, specify parameters like output format, and receive a structured response.
Learning in the Age of AI Assistants
AI assistants have fundamentally changed how we learn technical skills. You can now learn interactively — asking an AI assistant to explain a concept, write example code, debug errors in real time, and explain what went wrong. You don't need to memorise how to parse a JSON response or iterate through a list of files. You ask the AI assistant to write that code, then focus on understanding what it produced and whether it solves your problem correctly. The skill shifts from code production to code comprehension and system design.
Skill 3: Taking Control — Structured Outputs and Predictable AI Behaviour
The critical challenge with using AI systems is this: when you ask a language model to extract information from a clinical paper, what exactly do you get back?
Without structure, you get prose. The model might write "This randomised controlled trial enrolled 247 patients with type 2 diabetes, with the primary endpoint of HbA1c reduction at 12 weeks showing a mean difference of -0.8% (95% CI: -1.2 to -0.4, p=0.001)." That's excellent for reading, but terrible for systematic analysis. How do you parse that text programmatically to populate a spreadsheet? How do you ensure the model includes all the fields you need?
This is where structured outputs transform everything. Instead of accepting whatever format the AI chooses, you define the exact structure of the response — specific fields, required elements, data types, and validation rules. The model must return information matching your schema, or the API call fails. This single capability is what makes GenAI suitable for professional research workflows where consistency, completeness, and parsability matter.
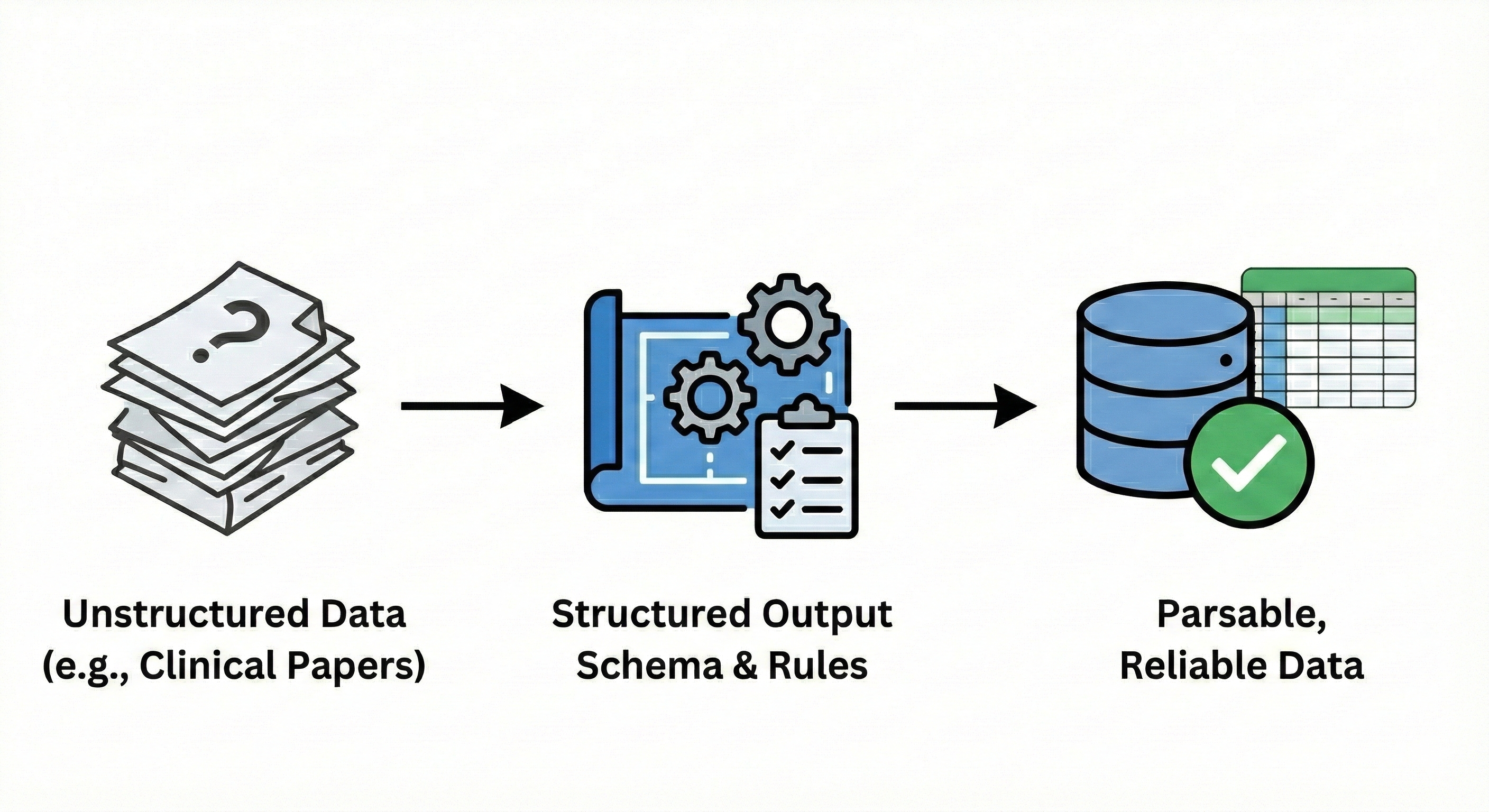
From Text Generation to Data Extraction
Consider the practical difference. Using a chatbot interface or unstructured API call, you might
prompt: "Extract key information from this clinical trial paper." The model returns a paragraph or
bullet points in whatever format it prefers. With structured outputs, you instead define a schema
like JSON — specifying exactly what data you want and in what format. For clinical trial extraction,
your schema might specify: study_design (string, required),
population_size (integer, required), primary_endpoint (string, required),
primary_endpoint_result (object with fields: metric, value, confidence_interval,
p_value), secondary_endpoints (array of objects), and adverse_events
(array of objects). The model returns a JSON object matching your specification completely, or
triggers an error.
Real-World Performance and Reliability
The practical impact is substantial. When two large language models (GPT-4-turbo and Claude-3-Opus) were used with structured extraction schemas to pull data from systematic review papers, concordance between models reached 87–96% depending on the complexity of the data being extracted. When the models produced concordant responses, accuracy was 0.94 — meaning structured extractions matched human expert review 94% of the time.
A systematic review workflow documented recently used structured outputs with GPT-4.1 for screening and specialised extraction schemas for data collection — researchers reproduced and updated an entire issue of Cochrane systematic reviews, 12 complete reviews, in under two days. The key enabler wasn't just AI capability, but the ability to define extraction schemas matching Cochrane's data collection requirements, ensuring every review captured the same standardised set of study characteristics, outcomes, and quality indicators in machine-parseable format.
The Practical Skill: Schema Definition
The actual skill you need to develop is schema definition — the ability to look at a research task and define what structured information you need extracted. This is less technical than it sounds, because it's fundamentally a domain expertise question dressed in technical clothing. Take economic modelling inputs as an example. You need treatment effectiveness data, cost parameters, utility values, and resource utilisation patterns from published literature. With structured outputs, you translate these domain requirements into schema format, defining fields for intervention name, comparator name, effectiveness metric, effectiveness value, effectiveness CI, cost year, currency, unit cost, resource use quantity, utility baseline, utility change, and so on. The JSON schema syntax has some technical elements, but AI assistants can write schema definitions from natural language descriptions of what you need.
Skill 4: Systems Thinking — Decomposing Complex HEOR Tasks
You've learned Python basics, gained API access, and mastered structured outputs. These skills give you control over individual AI operations — sending a document to a model, defining the response format, collecting structured data. But there's a fundamental problem we haven't yet confronted: the work you actually do in HEOR isn't captured by individual operations.
Systematic literature reviews don't consist of "extract data from one paper." They involve search strategy development, database queries, title/abstract screening across thousands of citations, full-text retrieval, eligibility assessment, structured data extraction, risk of bias evaluation, evidence synthesis, and reporting. Cost-effectiveness models don't emerge from "build me a model." They require identifying clinical pathways, extracting transition probabilities from literature, gathering cost and utility parameters, validating model structure against clinical reality, implementing calculations, conducting sensitivity analyses, and generating outputs suitable for regulatory submission.
No single AI call completes these tasks. The solution isn't more powerful AI — it's systems thinking: the ability to decompose complex work into components with clear inputs and outputs, identify which components are suitable for AI automation, and design the connections that make them function as a coherent whole.
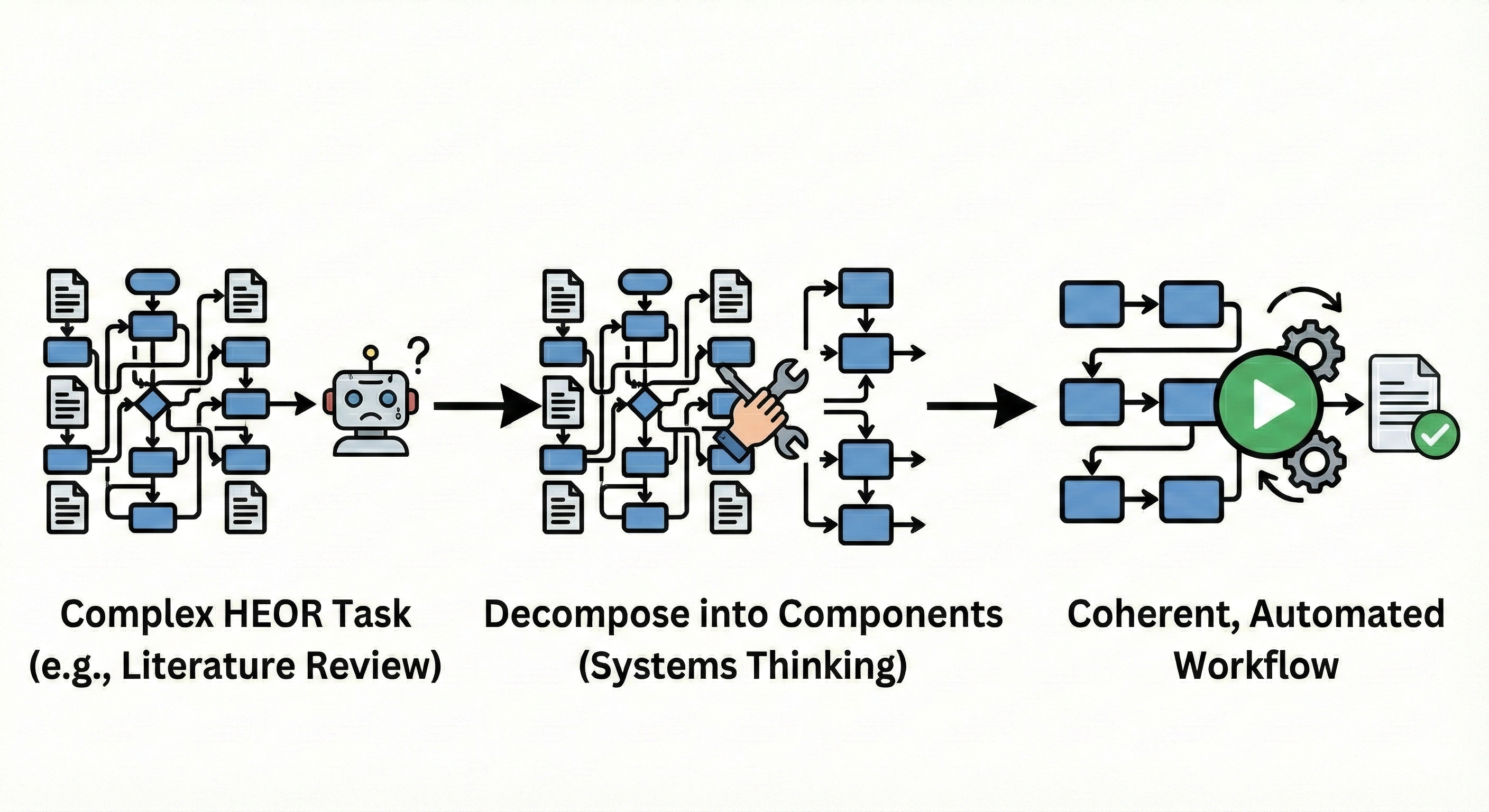
The Mental Model Shift That Matters
Instead of asking "What can AI do?", systems thinking asks "How do I decompose this research workflow into components, assign each to the best tool (AI or traditional code), and orchestrate them into a reliable pipeline?" This mental model unlocks ambitious applications. Consider a systematic literature review for a cost-effectiveness model. Decomposed properly:
- Title/abstract screening is a high-volume, pattern-recognition task — highly AI-suitable using structured outputs (include/exclude decision, confidence score, exclusion reason)
- Eligibility assessment involves collaborative AI–human workflow, where the model provides initial assessment, flags edge cases, and routes borderline studies to human reviewers
- Data extraction becomes a structured AI operation using the schemas we discussed — the model extracts predefined study characteristics, outcomes, and quality indicators into machine-parseable format
- Cross-validation employs multi-model checking, running extractions through both GPT-4 and Claude, automatically identifying discordances, and using model-critique workflows to resolve disagreements
- Evidence synthesis and reporting remains primarily human, with AI assisting in draft generation and formatting
Notice the pattern: the system succeeds not because AI does everything, but because each component plays to its strengths. AI excels at pattern matching (screening), structured information extraction, and text generation. Deterministic code handles file operations, API queries, and data validation. Humans provide judgment on methodology, interpret edge cases, and ensure clinical and scientific validity.
Building Connections: The Orchestration Layer
Decomposition identifies components. The orchestration layer connects them into functioning systems. This is where Python skills and API access combine with systems thinking to create something greater than the sum of parts. In practical terms, orchestration means writing code that: manages workflow state (which papers have been screened, which are awaiting extraction, which need human review); routes information between components (screening outputs become extraction inputs); handles errors gracefully (what happens when PDF retrieval fails or extraction returns invalid data); implements quality gates (automated validation before proceeding to next step); and maintains audit trails (logging what happened at each stage for reproducibility and compliance).
Research teams building production AI systems for systematic reviews report that the orchestration code — the "glue" connecting components — often represents 30–40% of total development effort, more than any single component. This isn't wasted effort. It's what makes the system reliable, maintainable, and trustworthy for professional research.
Skill 5: Building Trust — Monitoring and Reliability in AI Systems
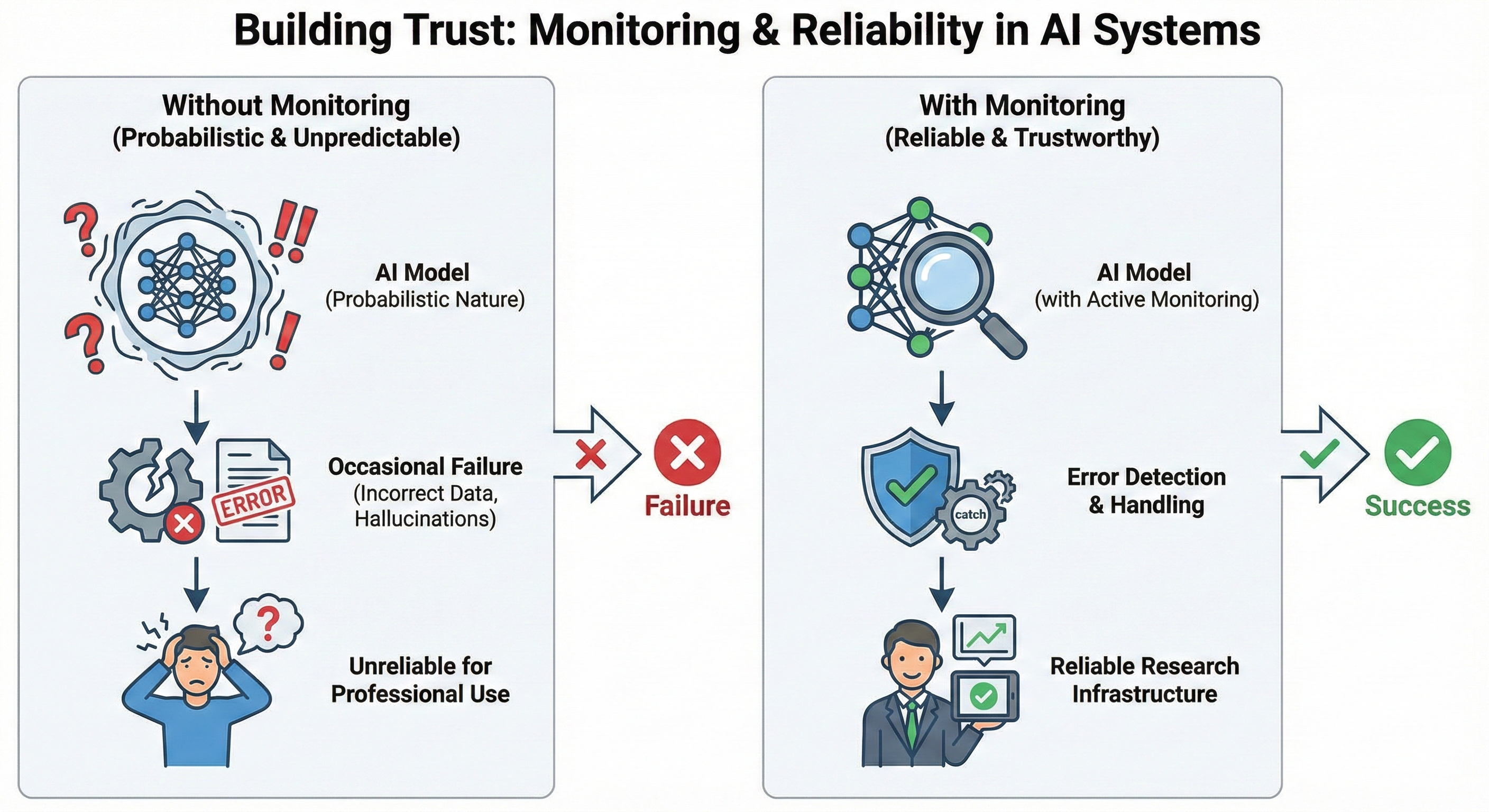
The final skill addresses a fundamental reality of working with probabilistic systems: unlike deterministic software, the same input can produce different outputs at different times. For research applications, this isn't acceptable without appropriate oversight systems in place.
What Monitoring Actually Means
In traditional software, monitoring is relatively straightforward: you log whether operations succeeded or failed. Did the database query execute? Did the API return a response? Did the file write complete? These are binary outcomes with clear success criteria.
LLM monitoring is fundamentally different because the core operation — generating text based on learned patterns — doesn't always have binary success. The model always returns something. The question is whether what it returned is accurate, complete, appropriate, and useful. That requires semantic evaluation, not just technical logging.
Effective monitoring for AI systems means observability at multiple levels:
- Input/output logging: capturing exactly what prompt and data were sent to each model call, what parameters were used (temperature, model version, token limits), and what structured output was received. This creates an audit trail that enables reproduction and debugging. When an extraction seems wrong, you need to see precisely what the model was given and what it produced.
- Validation gates: automated checks that outputs conform to expected patterns and business rules. For clinical data extraction: verifying that extracted JSON matches your schema (technical validation), that required fields are populated (completeness validation), that numeric values fall within plausible ranges (sanity checking), and that relationships between fields make sense (logical validation).
- Performance metrics tracking: measuring accuracy, concordance, and reliability over time. When using multi-model collaborative extraction approaches — where GPT-4 and Claude independently extract data and you compare results — concordance rate becomes a real-time quality indicator. If concordance drops below your threshold for specific study types, your monitoring system routes those extractions to human review.
Building Reliable Systems in Probabilistic Environments
The goal of monitoring isn't eliminating AI errors — it's detecting them systematically and handling them appropriately. This changes how you design systems. Rather than hoping outputs are correct, you build validation at each step. Rather than manual spot-checking, you implement automated quality gates that flag issues without requiring human intervention for every output. Rather than discovering failures when results are already in a report, you catch them while they can still be corrected.
This is the foundation of professional practice with AI: not trusting individual outputs, but building systems that are trustworthy because their quality controls work reliably.
Your Weekend Starting Point: From Reading to Doing
The five skills described here aren't abstract recommendations. They're the specific capabilities that separate professionals who use AI as an occasional helper from those who have integrated it as a reliable research tool. Each is learnable. None requires becoming a software engineer or abandoning the domain expertise that defines your professional value.
Your Action This Weekend
Pick one real task from your current work — not a toy example designed to be easy, but something you actually need to accomplish. Perhaps you're conducting a literature review and facing 200 papers for data extraction. Maybe you're building an economic model and need to extract cost and utility parameters from 50 publications. Whatever the task, apply one skill to that task this weekend.
When you watch your Python script successfully extract treatment efficacy data from 20 papers in three minutes — work that would have taken you a full day manually — you understand the potential in ways no article can convey.
Building Momentum Through Compound Returns
The counterintuitive reality is that starting small doesn't mean achieving small results. These skills combine and multiply their impact: learning Python basics enables API access; API access enables structured outputs; structured outputs enable systems thinking; systems thinking requires monitoring; and monitoring informs improvement. Research teams consistently report moving from first API call to production systematic review workflows processing hundreds of papers in 8–12 weeks. The bottleneck isn't technical complexity once fundamentals are established — it's domain expertise about what workflows matter and how to validate outputs appropriately. That's precisely where HEOR professionals have an inherent advantage.
The path forward is clear: pick one skill, apply it to one real task this weekend, and experience what becomes possible when you move from reading about GenAI potential to building systems that deliver it. The barrier isn't capability — it's the decision to start. Make that decision. The compound returns are waiting.
Download the Full Paper
Get the PDF version with complete references for offline reading and sharing with your team.
Developed by Aide Solutions LLC. This white paper was prepared with the support of generative artificial intelligence tools. The author reviewed, edited, and takes full responsibility for the content and conclusions presented. Full references are available in the PDF version.